Phylogeny (amino acid)
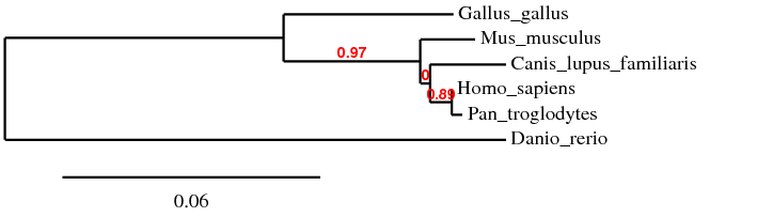
In this tree above, using protein sequences in the MABL program, there was little deviation to what is normally thought as the tree using other molecular markers or anatomical data. However, there is a bootstrap of 0 on the branch off of the mouse and dog lineages. This implies that there really no need for the extra branch, which should be condensed into the mouse branch, implying that mouse and dog Sox9 have the same amount of divergence from each other compared to the primate lineage. It is not surprising that zebrafish are the outgroup in this phylogram.
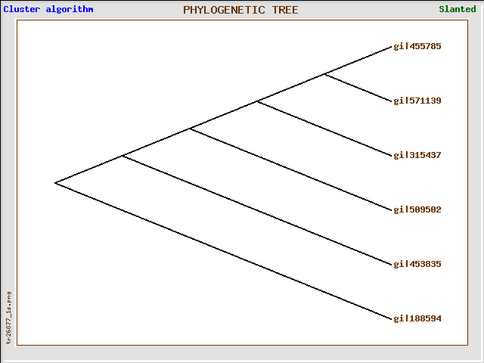
The tree generated from Gene Bee is essentially the same as the tree generated from MABL. However, I did not bootstrap this tree, hence the confidence in this tree is a bit shaky for me. I definitely prefer using MABL because MABL allows for an easy way to name your organisms represented, while I could not figure out how to replace the gi numbers with the names in the Gene Bee tree.